研究成果与卓越性 Research Excellence
在顶级期刊发表论文 (Nature、Nature Genetics、Cell Reports) Published in top-tier journals (Nature, Nature Genetics, and Cell Reports)
研究团队建立了Geo-seq、Auto-seq等单细胞空间组学技术平台 Research team developed Geo-seq, Auto-seq and other single-cell spatial omics platforms
利用单细胞和空间多组学技术研究细胞命运调控,发表 55 篇论文他引 1,705 次 Pioneering single-cell and spatial multi-omics for cell fate regulation, 55 publications with 1,705 citations
胚胎发育 4D 时空模型, 推断时间-空间-细胞命运的耦合调控 4D spatiotemporal embryonic development model, inferring coupled time-space-cell fate regulation
学术发表 Academic Publications
团队亮点 Group Highlights
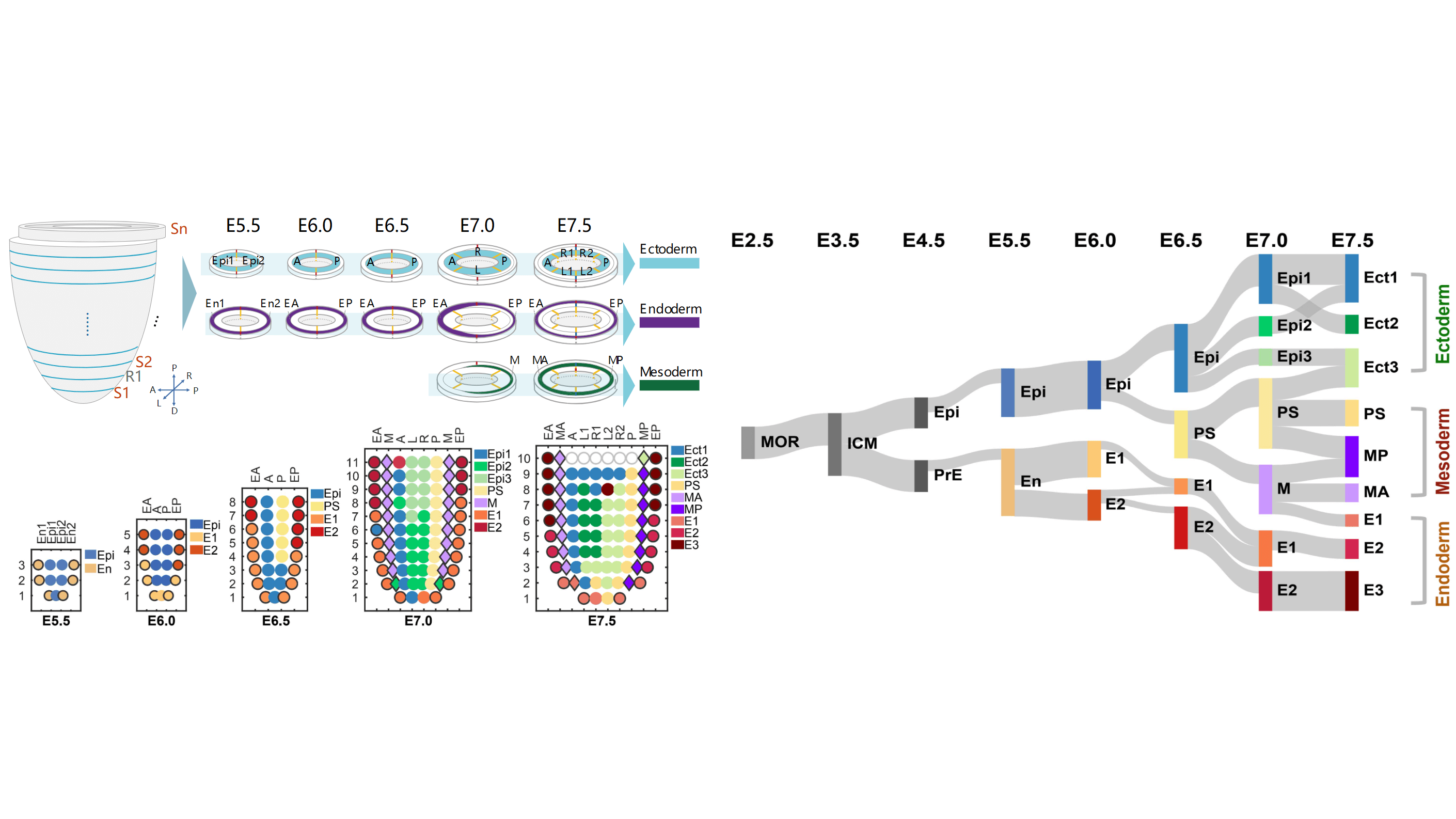
Guizhong Cui#, Su Feng#, Yaping Yan#, Li Wang, Xiechao He, Xi Li, Yanchao Duan, Jun Chen, Patrick P.L. Tam, Ke Tang, Ping Zheng, Wei Si+, Naihe Jing+, Guangdun Peng+. Spatial molecular anatomy of germ layers in the gastrulating Cynomolgus monkey embryo. Cell Reports, 2022, 40(9): 111285-111285.
Guangdun Peng#+, Shengbao Suo#, Guizhong Cui#, Fang Yu#, Ran Wang, Jun Chen, Shirui Chen, Zhiwen Liu, Guoyu Chen, Yun Qian, Patrick P. L. Tam, Jing-Dong J. Han+, Naihe Jing+. Molecular architecture of lineage allocation and tissue organization in early mouse embry. Nature, 2019, 572(7770): 528-32.
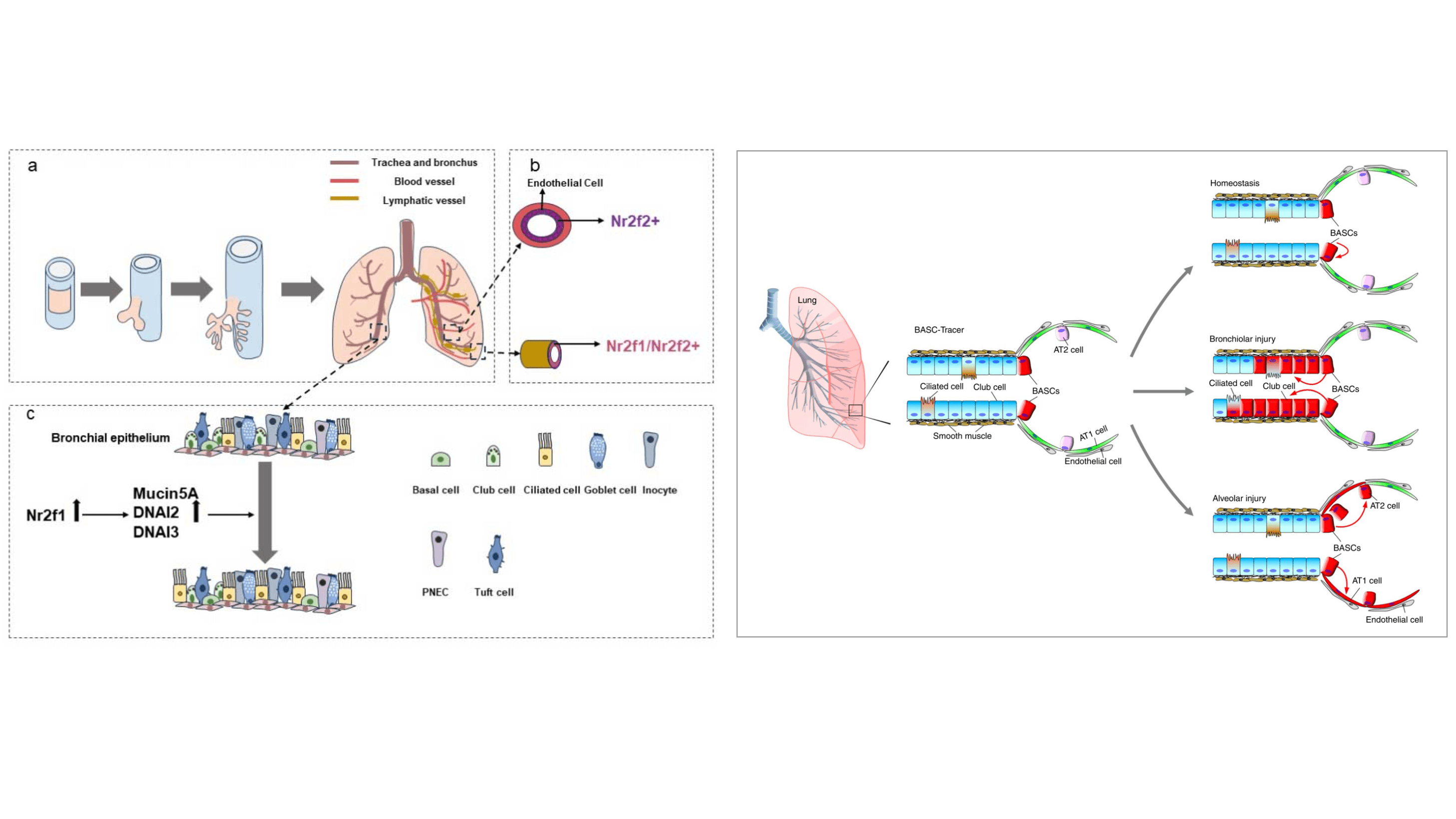
Qiaozhen Liu#, Kuo Liu#, Guizhong Cui#, Xiuzhen Huang, Shun Yao, Wenke Guo, Zhen Qin, Yan Li, Rui Yang, Wenjuan Pu, Libo Zhang, Lingjuan He, Huan Zhao, Wei Yu, Muxue Tang, Xueying Tian, Dongqing Cai, Yu Nie, Shengshou Hu, Tao Ren, Zengyong Qiao, Hefeng Huang, Yi Arial Zeng, Naihe Jing, Guangdun Peng+, Hongbin Ji+, Bin Zhou+. Lung regeneration by multipotent stem cells residing at the bronchioalveolar-duct junction. Nature Genetics, 2019, 51(4): 728-38.
Jiaxin Yang, Wenjing Sun, Guizhong Cui+. Roles of the NR2F Family in the Development, Disease, and Cancer of the Lung. Journal of Developmental Biology, 2024, 12(3): 24.
第一作者/通讯作者 First Author/Corresponding Author
1. Xiaogao Meng#, Wenjia Li#, Jian Xu, Yao Yao, An Gong, Yumeng Yang , Fangfang Qu, Chenkai Guo, Hui Zheng, Guizhong Cui+, Shengbao Suo+, Guangdun Peng+. Spatio-temporal transcriptome atlas of developing mouse lung. Science Bulletin, 2025, 70(10): 1641-58.
2. Jiaxin Yang, Wenjing Sun, Guizhong Cui+. Roles of the NR2F Family in the Development, Disease, and Cancer of the Lung. Journal of Developmental Biology, 2024, 12(3): 24.
3. Fangfang Qu#, Wenjia Li#, Jian Xu#, Ruifang Zhang, Jincan Ke, Xiaodie Ren, Xiaogao Meng, Lexin Qin, Jingna Zhang, Fangru Lu, Xin Zhou, Xi Luo, Zhen Zhang, Minhan Wang, Guangming Wu, Duanqing Pei, Jiekai Chen, Guizhong Cui+, Shengbao Suo+, Guangdun Peng+. Three-dimensional molecular architecture of mouse organogenesis. Nature Communications, 2023, 14(1): 4599.
4. Xiaogao Meng, Guizhong Cui+, Guangdun Peng+. Lung development and regeneration: newly defined cell types and progenitor status. Cell Regeneration, 2023, 12(1): 5.
空间转录组学数据库 Spatial Transcriptomics Database
Spatial Of Primate (SOP)
猕猴胚胎发育空间转录组学数据库 Cynomolgus Monkey Embryogenesis Spatial Transcriptomics
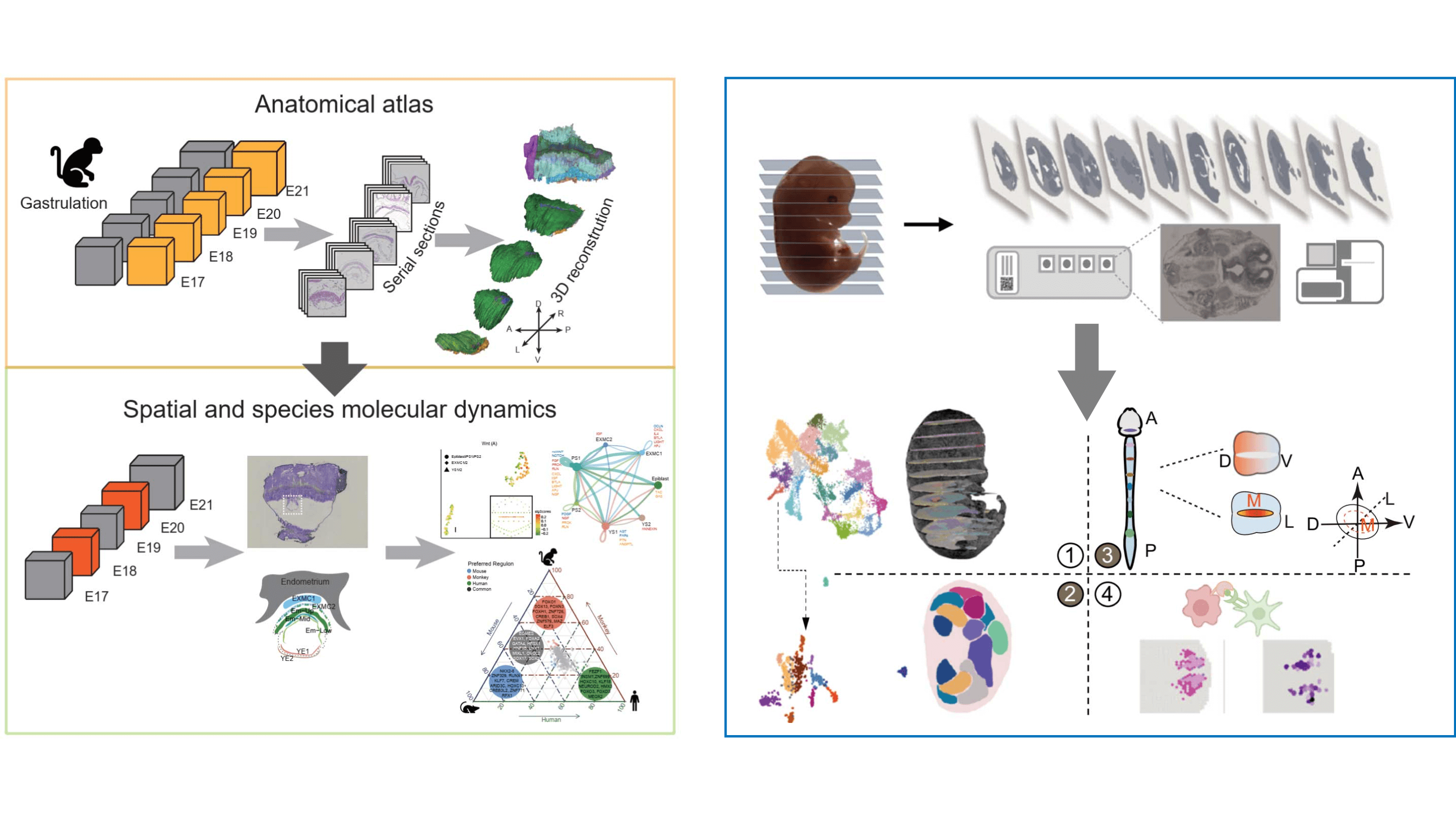
基于Geo-seq技术构建的猕猴胚胎发育高分辨率解剖学图谱,提供3D数字胚胎模板和空间转录组学数据,支持跨物种比较分析。 High-resolution anatomical atlas of Cynomolgus gastrulation embryos based on Geo-seq technology, providing 3D digital embryo templates and spatial transcriptomics data for cross-species comparative analysis.
3D重建 3D Reconstruction
E17-E21发育阶段3D模型 E17-E21 stage 3D models
空间转录组学 Spatial Transcriptomics
胚层特异性基因表达 Germ layer-specific expression
跨物种比较 Cross-species Analysis
小鼠与灵长类对比 Mouse vs Primate comparison
数据下载 Data Download
原始数据和分析结果 Raw data & analysis
技术亮点 Technical Highlights
- • 基于Geo-seq技术的空间转录组学Geo-seq based spatial transcriptomics
- • 1天时间分辨率的发育动态1-day temporal resolution dynamics
- • 连续切片重建的3D数字胚胎Serial sections 3D digital embryo
- • 交互式可视化平台Interactive visualization platform
科学发现 Scientific Insights
- • 胚层形成的进化保守性Conserved germ layer formation
- • 物种特异性转录程序Species-specific programs
- • 发育坐标系的建立Developmental coordinate system
- • 信号通路的空间调控Spatial signaling regulation